Scientific Reports (2020)
Julian Wolf, Claudia Auw-Haedrich, Anja Schlecht, Stefaniya Boneva, Hans Mittelviefhaus, Thabo Lapp, Hansjürgen Agostini, Thomas Reinhard, Günther Schlunck, Clemens Lange
Number of citations (crossref.org): Loading....

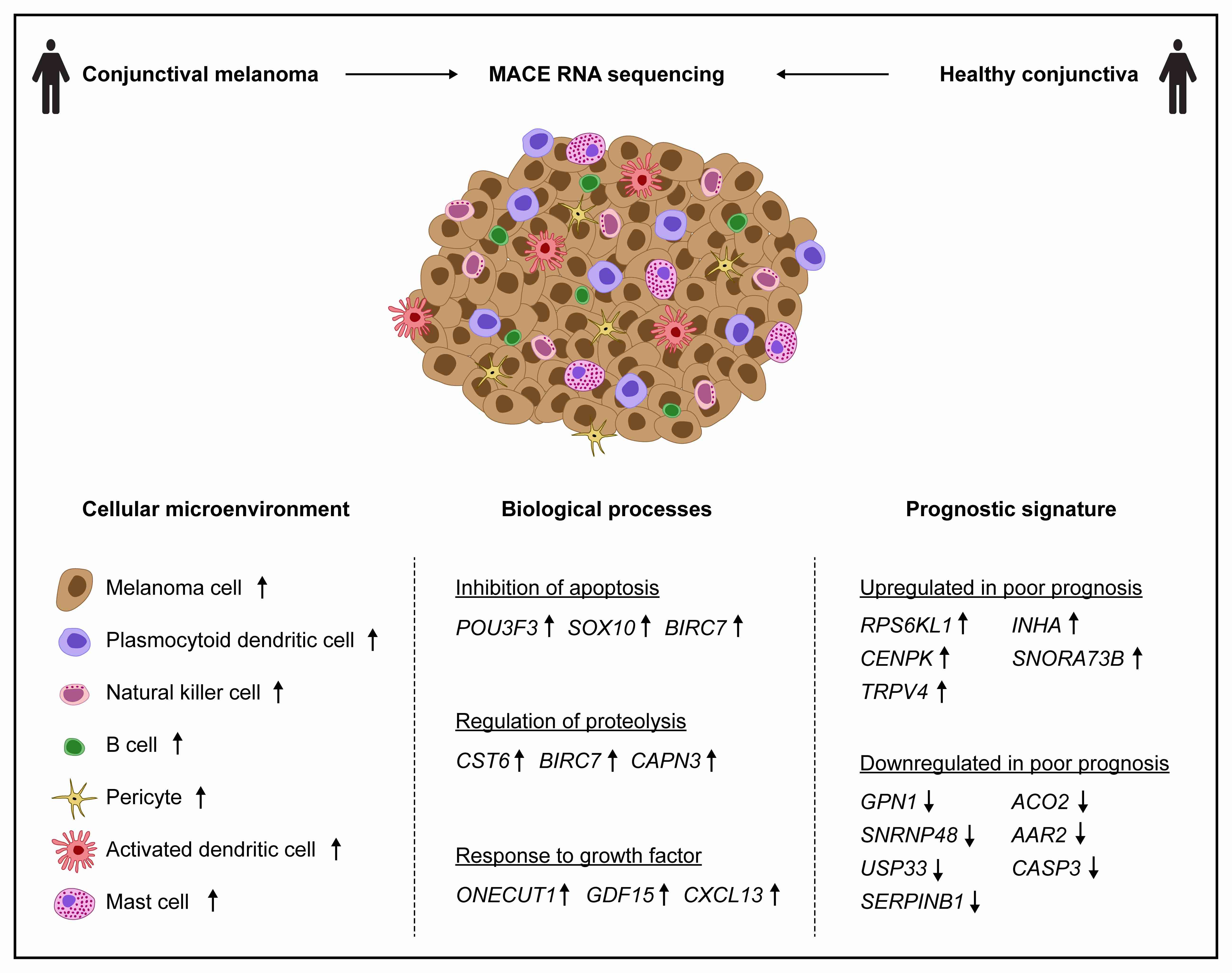
This study characterizes the transcriptome and the cellular tumor microenvironment (TME) of conjunctival melanoma (CM) and identifies prognostically relevant biomarkers. 12 formalin-fixed and paraffin-embedded CM were analyzed by MACE RNA sequencing, including six cases each with good or poor clinical out-come, the latter being defined by local recurrence and/or systemic metastases. Eight healthy conjunctival specimens served as controls. The TME of CM, as de-termined by bioinformatic cell type enrichment analysis, was characterized by the enrichment of melanocytes, pericytes and especially various immune cell types, such as plasmacytoid dendritic cells, natural killer T cells, B cells and mast cells. Differentially expressed genes between CM and control were mainly involved in inhibition of apoptosis, proteolysis and response to growth factors. POU3F3, BIRC5 and 7 were among the top expressed genes associated with inhibition of apoptosis. 20 genes, among them CENPK, INHA, USP33, CASP3, SNORA73B, AAR2, SNRNP48 and GPN1, were identified as prognostically relevant factors reaching high classification accuracy (area under the curve: 1.0). The present study provides new insights into the TME and the transcriptional profile of CM and additionally identifies new prognostic biomarkers. These results add new diagnostic tools and may lead to new options of targeted therapy for CM.